k-means
An unsupervised learning algorithm that partitions data points into clusters by iteratively assigning points to the nearest cluster center and updating centers as the mean of assigned points.
Given data and number of clusters , it minimizes:
where if point belongs to cluster (else 0), and is the centroid of cluster .
First we initialize centroids (often randomly from data points)
The algorithm alternates between two steps until convergence:
- Assignment: Assign each point to the nearest centroid: for each point : assign to cluster
- Update: Recompute centroids as the mean of assigned points: for each cluster : update
where is the set of points assigned to cluster .
This simple procedure guarantees that the objective decreases monotonically, though it only finds local minima.
The final clustering depends heavily on initialization - poor starting centroids can lead to suboptimal solutions.
Practical considerations / Properties & limitations
Choosing : Often determined by domain knowledge or methods like the elbow method (plot reconstruction error vs ) or silhouette analysis.
Initialization: k-means++ improves on random initialization by choosing centroids that are far apart, leading to better convergence and final clusters.
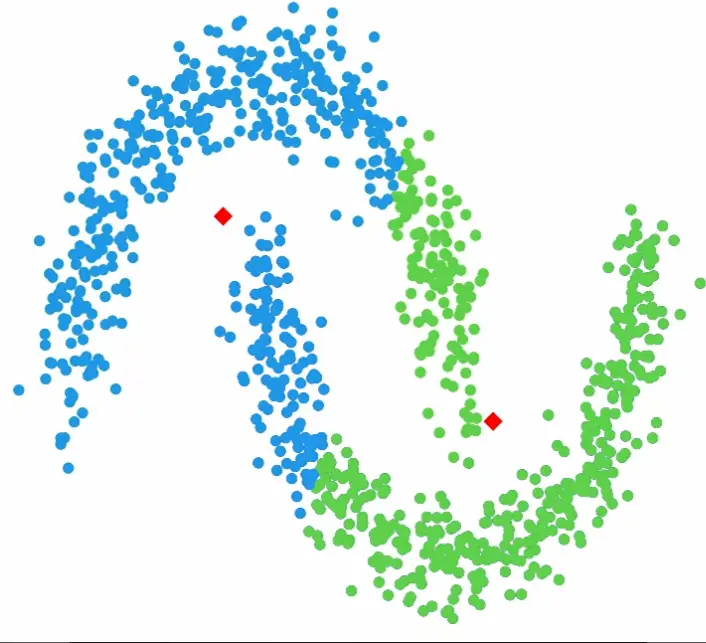
Assumptions: k-means implicitly assumes spherical clusters of similar size. It struggles with elongated or overlapping clusters, where algorithms like gaussian mixture models or DBSCAN may perform better.
K-means focuses on minimizing variance within clusters (“compactness” within clusters) but does not explicitly maximize separation between clusters.Sensitive to noise an outliers: Outliers can skew centroids significantly. Preprocessing steps like outlier removal or using k-medoids can help.
The algorithm’s complexity per iteration (for -dimensional data) makes it efficient for large datasets. Variants include mini-batch k-means for even larger data and kernel k-means for non-linearly separable clusters.
Example data it struggles with: